Reading signals from cultured neurons
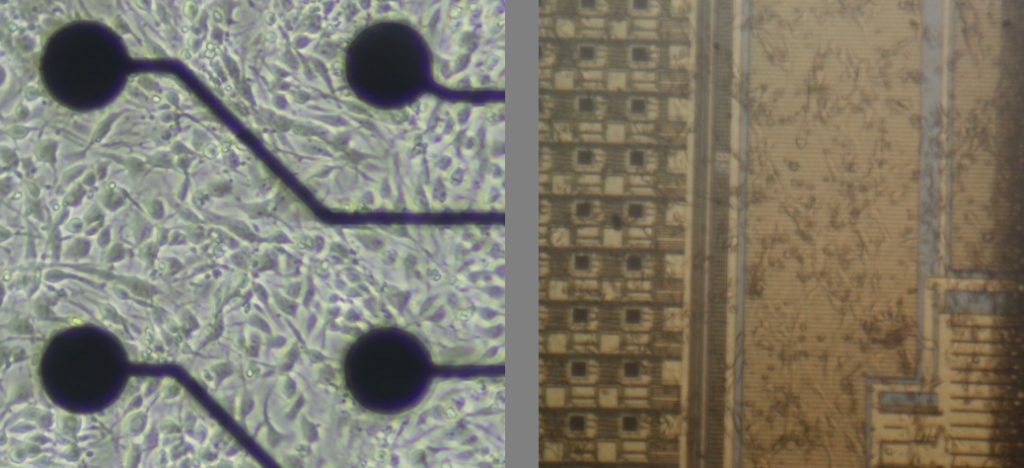
Figure 1. Low density vs high density MEAs interfaced to in vitro neural cell cultures.
From a system automation point of view, a Biological Neural Network (BNN) unit may be seen as an analogue system which can be read, for various analysis and monitoring purposes, through sensors. It may also be commanded (written to) by a diversity of stimulation mechanisms, either chemical, electrical, or even optical. For in vitro BNNs cultures, recent electrophysiology technologies facilitate such a setup, as multi-electrode arrays (MEA) integrated devices enable a direct interface from multiple parallel channel readouts intertwined with electrical stimulation probes in contact to the neural cell culture directly grown over the MEA electronic circuit.
First generation passive electrophysiology probes lack spatial accuracy: they only enable readouts at low scale range, from a group of cells rather than at a single-neuron level. These technologies nevertheless provide certain insights on the neural network activity. Low-scale MEA technology has thus become popular throughout academic and industrial laboratories worldwide in the past couple of decades, to the extent that commercial systems start to get commoditized, in particular for pharmacological and toxicity assay applications [1][2].
Thanks to further integration using CMOS technology in the past decade, it has become possible to increase both the spatial density (up to several thousands of readout electrodes and dozens of stimulation electrodes distributed over a few square millimeters neuron culture surface area) and temporal resolution (up to sub-millisecond accuracy) of BNN systems [3]. CMOS circuits also facilitate the generation of spatiotemporal signal patterns for stimulation probes, thus paving the way for the next generation experiments we anticipate in closed-loop operations (e.g. for artificial training). These integrated devices typically embed amplifiers and analogue-to-digital signal conversion, so that they can be directly interfaced to digital networks such as USB or Ethernet interfaces, digital memories and personal computing devices [4]. Depending on the needs of the target applications, the MEA signals may be recorded and analyzed at different scales, ranging from low resolution network activity sampling (e.g. spiking activity) to high-resolution measurements at the sub-cellular level (e.g. axonal action potential propagation) [5]. The amount of readout data they generate at high-resolution is however tremendous, thus requiring dedicated signal compression, multiplexing and filtering on-chip processing and software tools [6]. In search for a trade-off between active on-chip sub-millisecond spike recording at high signal to noise ratios and large-area recordings at higher spatial scales with more conventional passive MEA probes, Angotzi et al. recently developed a multi-scale recording system combining different technologies [7].
Scaling-up BNN architectures therefore requires multidisciplinary engineering approaches to integrate the wetware constraints (growing and maintaining the neural cell culture), the hardware constraints (interfacing and operating sensors and actuators to the neural cell culture with as little disruption as possible) and software constraints (signal analysis and signal synthesis at high resolutions, both spatial and temporal) into operational, sustainable and reliable systems. After our initial period of incubation at the Biopole Startlab in Lausanne, Switzerland to train our engineering team on wetlab protocols and preliminary experiments with MEA technologies, we are now ready to move to the next stage at our own laboratory infrastructure in Vevey. Stay tuned!
References
- Neuroservice Corporate presentation slides, 2015 – https://www.slideshare.net/BrunoBuisson/neuroservicecorporate2015
- Neuroproof and Fujifilm Cellular Dynamics Webinar, 2017 – https://www.youtube.com/watch?v=6Ol_aW_Pajs
- Kim R. et al., “Recent Trends in Microelectrode Array Technology for In Vitro Neural Interface Platform”, Biomed Eng Lett (2014) 4:129-141
- Obien M.E.J. et al., « Revealing neuronal function through microelectrode array recordings”, Frontiers in Neuroscience, Systems Biology, Vol.8, Jan. 2015.
- Muller J. et al., “High-resolution CMOS MEA platform to study neurons at subcellular, cellular and network levels”, Lab Chip 15 2067, 2015.
- Maccione A. et al, “Microelectronics, bioinformatics and neurocomputation for massive neuronal recordings in brain circuits with large scale multielectrode array probes”, Brain Research Bulletin, Vol.119, Part B, pp. 118-126, Oct. 2015.
- Angotzi G.N. et al., « A Synchronous Neural Recording Platform for Multiple High-Resolution CMOS Probes and Passive Electrode Arrays”. IEEE Transactions on biomedical circuits and systems, Vol. 12, No. 3, June 2018.